Zebras, not horses: The genetic disease that can’t be sequenced

Growing up, I was a weird bendy kid. I used to run, dance, do gymnastics, cycle, and was uncontrollably energetic and fidgety. When I was 11 years old, my ankle slipped out of place. It was bright purple, puffy, and stopped me from dancing. When I was 13, my shoulder started dislocating every time I raised my arms to put a t-shirt on. By the age of 16, my fingers, toes, and wrists were clicking and cracking. By the age of 21 I was diagnosed with a heritable genetic disease called Ehlers Danlos Syndrome, or EDS. I was told I have the most common type, hypermobile EDS, but there was no genetic marker for it. It took me 10 years to be diagnosed, and it was only after I’d begun a Bachelor’s degree in genetics that I was taken seriously. What took me so long to be diagnosed?
Most of us have heard of Occam’s razor, and may have been told: “When you see hoof prints, think horses, not zebras”. It’s an age-old parable to remind trainee doctors that the simplest cause is most often the answer. For 10 years, I was called a hypochondriac, told I had an anxiety disorder, that it was growing pains. Sometimes, the easiest explanation is not always the correct diagnosis.
The Ehlers Danlos
EDS was first discovered in the early 20th
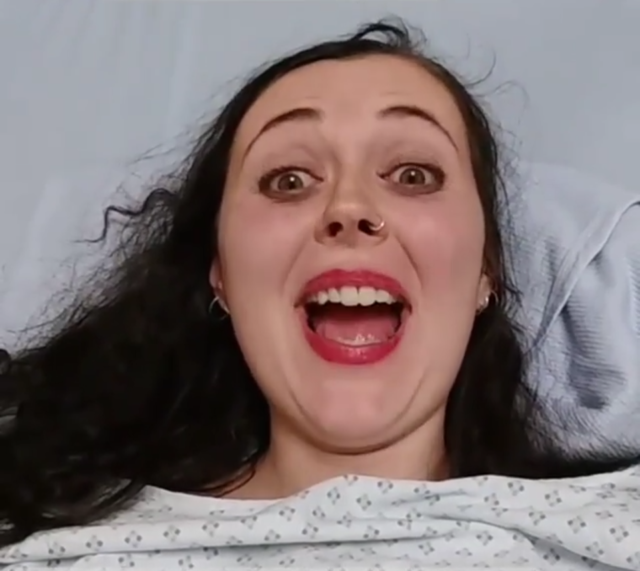
Me, on the highest possible morphine dose in Glasgow’s Royal Infirmary after dislocating my entire left leg in 2017
Understandably, this causes a host of problems for EDS sufferers: easy bruising, dislocations, partial dislocations, digestive problems, severe fatigue, abnormal scarring,
The struggle to diagnose Ehlers Danlos syndromes
The first hurdle is acknowledging that sometimes it might be zebras and not horses causing the symptoms. The Ehlers Danlos Foundation recommends that “If you can’t connect the issues, think connective tissues”, and this moniker has been relatively fruitful in encouraging health care professionals to assess patients with seemingly unconnectable issues3. There are 13 types of EDS, some of which only have a handful of cases reported globally.
Luckily, 12/13 types of EDS have distinct and
How do you sequence a gene?
Normally, if a specific disease is expected, a DNA swab from blood, hair, or saliva is taken from the patient. The known, or suspected, disease-causing regions are amplified so there’s enough DNA to work with. From there, the DNA can be sequenced. This involves detecting the order of adenosine, cytosine, guanine, and thymine (the four DNA bases) in the DNA. If the bases are in the wrong order, for example, if one base is swapped for another this can cause a mutation, leading to a faulty version of the gene and the protein it codes for. Sometimes, entire genes are deleted too. In the 12/13 EDS variants that are
A needle in a haystack: finding the cause
Why hasn’t the causative gene been identified yet? There are quite a few theories floating around. Unfortunately, the main reason is possibly due to financial constraints: we’re looking for a genetic needle in a global haystack. Luckily enough, a very rich mysterious benefactor (no, seriously), has donated £1,000,000 towards the sequencing efforts. This will involve sequencing the entire genomes of 1000 EDS hypermobile type sufferers globally, cross-comparing gene mutations, and hopefully finding the cause6.

Or, it may be possible that the disease is not caused by a structural deficiency in collagen at all. In 2016, a group of researchers discovered a gene expansion
The outlook for people with EDS can be vastly different. Whilst not life-threatening, hypermobile EDS can severely compromise the quality of life for a number of people. I’m lucky to (just about) be walking around on my own two feet, using a crutch occasionally. Others are
If you are diagnosed with Ehlers Danlos Hypermobility type, consider signing up for the 1000 genomes project, here.
This article was specialist edited by Kirstin Leslie and copy-edited by Caitlin Duncan.
References
- https://www.ncbi.nlm.nih.gov/pubmed/29446032
- https://www.mdedge.com/psychiatry/article/161887/anxiety-disorders/anxiety-and-joint-hypermobility-unexpected-association
- https://ehlers-danlos.com/wp-content/uploads/hEDS-Dx-Criteria-checklist-1.pdf
- https://www.ncbi.nlm.nih.gov/books/NBK1279/
- Check this reference here for a more detailed explanation http://www.mastcelldisease.com/mcas-ehlers-danlos-syndrome-eds-exploring-connection/
- https://www.ehlers-danlos.com/million-pound-gift/
- https://www.niaid.nih.gov/research/hereditary-alpha-tryptasemia-faq
- https://www.mdedge.com/rheumatology/article/115772/dermatology/tryptase-gene-variant-linked-gi-joint-and-skin-symptoms