Differences in Gene Regulation Between Mice and Humans
Mammalians have around 25,000 genes, but only 62% of the human genome is transcribed in one or more cell types 1. Considering this, it’s possible to understand that the performance of a cell is not only determined by its gene set, but rather by which genes are active, which is highly variable among cells. All cells within an organism are genetically identical, but in each cell only part of the genes are expressed, namely in a unique gene expression pattern, allowing the cell to perform its function. Gene activity is regulated by transcription factors that bind to regulatory sequences in relative proximity to the gene. For this purpose, every gene has a promoter sequence which is essential for gene expression. Mouse and human genes are fairly conserved, as most genes have a corresponding gene in the other species. However, a better understanding of the differences between the human and mouse genome is very important, as mice are widely used as disease models to better understand and treat human diseases.
Recently, three papers detailing the ways that mouse and human genomes are regulated appeared in the journal Nature. They investigated transcription factor binding patterns across the genomes and DNA sequences that potentially bind transcription factors. Among the main findings is the discovery that the regulatory patterns between different tissues of the same species are more similar than between the same tissues in mouse and human, meaning that the regulatory pattern in my heart is more similar to the one in my kidney than to the one in the heart of a mouse 2. This fact may raise the concern that the mouse is too different from us to be studied to understand human diseases. Another major finding was that transcription factors binding to promoter regions is highly conserved, whereas binding to more distant regulatory elements is more lineage specific 3.This suggests that the rough gene regulatory pattern is conserved, whereas the fine tuning is rather species specific.
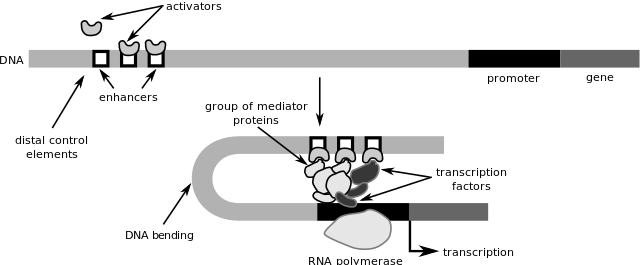
Transcription factor binding to enhancer and promoter regions regulates gene activity. Image credit: Philippe Hupé via Wikimedia Commons (License)
Looking at specific cellular processes, the authors found that genes involved in intracellular processes such as RNA processing, chromatin organization and several intracellular metabolic processes tend to have a similar gene expression pattern between human and mouse, whereas genes involved in extracellular matrix, cellular adhesion and signalling receptors are less conserved. Most importantly, results showed that regulatory sequences next to immune-system-related genes are very divergent. This may suggest that gene expression profiles of basic cellular functions are well conserved, whereas gene regulation for cell-cell interactions and the organisation between cells within an organism is more diverged.
In respect of the importance as a disease model, the researchers found that many gene variants that are associated with a certain disease, for instance liver genes that are related to cholesterol and alcohol dependence, can also be found in the mouse genome. These findings are very useful in respect of using the mouse as a disease model and will facilitate with regards to which diseases mouse models are appropriate for.
Edited by Debbie Nicol
References
- Yue, F. et al, A comparative encyclopedia of DNA elements in the mouse genome. Nature 515, 355-364 (2014)
- Stergachis, A. B. et al, Conservation of trans-acting circuitry during mammalian regulatory evolution. Nature 515, 365-370 (2014)
- Cheng, Y. et al, Principles of regulatory information conservation between mouse and human. Nature 515, 371-375 (2014)










